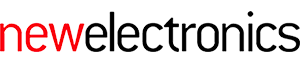Wilder Penfield’s experiments in the early 1950s with electrical neural stimulation helped map areas of the brain for the first time. The homunculus who appears in biology textbooks, showing which parts of the brain are responsible for touch, smell, taste and movement, was the result of this work.
Although these early experiments helped elucidate much about the way in which the brain works, the methods for interfacing electronics to biology have not advanced quickly. A major problem is that electrical stimulation and recording from outside the cells is a blunt instrument for picking up what is happening at a cellular level. In the watery environment of the body, electrical stimulus can easily interfere with multiple cells at a time and sensors get an extremely blurred picture of what is happening to the cells around them.
Focused or laser light can provide a more precise way to stimulate cells and also to read data from them. Potentially light can also show changes in activity not just per cell but as signals move through the cell and onto to neighbouring synapses. But it has a major drawback: it only works on cells that have been genetically modified to produce and react to light. Used on cells in culture and modified test animals, optogenetics proved invaluable for scientists who want to see how neurons in culture communicate with each other. Recent advances in identifying light-emitting proteins, such as a project at MIT led by associate professor Edward Boyden that analysed 1.5 million variants of one promising proteins, is helping to increase the resolution of the photon signalling to track changes within neurons and synapses.
When it comes to dealing with live patients, electrical stimulation and measurement remains the primary mechanism and, despite the low resolution, has led to the point where brain-computer interfaces let people with severe spinal injuries gain control of artificial limbs. They may, in time, provide a way to bridge severed connections and reconnect organic limbs to the brain’s signals. One of the reasons why brain-computer interfaces work is that the brain itself adapts to their presence. Through lengthy training, patients can direct activity to the array of neurons closest to the probes that result in useful movement. But the technology remains some way from being able to decode brain activity directly without relying on the ability to train the user.
 Above: Columbia's sensor technology builds ion-channel proteins into an artificial lipid membrane contained within the package
Above: Columbia's sensor technology builds ion-channel proteins into an artificial lipid membrane contained within the package
Integrated silicon sensors
In terms of probe resolution, highly integrated silicon sensors with multiple electrodes across their surface is increasing precision.
Research institute Imec made its Neuropixels probe available to neuroscientists late last year. The probe has 384 dual-band recording channels that can be picked from almost a thousand conductive electrodes tiled across the 10mm-long flat, square tip. In principle, the probe will make it possible to record signals from hundreds of neurons that touch its surface and is light enough to support experiments with freely moving animals.
For the kind of resolution needed to detect neuronal activity in humans, light may still provide an answer. One method is to employ gene therapy on the recipient, though this comes with serious ethical challenges. Neurons in the vicinity of an injection can be infected using a carrier virus that inserts the genes used to create light-emitting proteins. An alternative is to employ infrared light to stimulate neurons, a technique discovered by scientists at Vanderbilt University at around the same time that optogenetics became a practical tool for research.
As well as helping to improve on the resolution of electrical measurement, the technique offers enough accuracy to map nerve impulses with applications in neurosurgery to help guide incisions when removing tumours as well as connecting prostheses to nerve endings. The main challenge is to find a way to miniaturise laser arrays to the point where they can interact with groups of neurons in a living brain.
In the past few years, sound has emerged as another potential way to stimulate and suppress neuron activity. Experiments showed it was possible decades ago, but they used damagingly high levels of ultrasound. William Tyler of Arizona State University working with colleagues at Johns Hopkins found low-intensity ultrasound can control neurons and map their activity to reasonably high precision with less risk of damage, mainly through self-heating, although the scientists caution more work is needed to test the systems. A key potential advantage of the technique is that the probes need not penetrate the skull; they can lie on the surface of the skull. But there are a number of obstacles to overcome to get the required level of focusing from speaker arrays and to work out whether the technique is, indeed, safe.
Working like cells
Ken Shepard’s group at Columbia University is taking a different approach to bio-interfacing by extending the usable signals beyond light or electrical impulse. It is one that could be used with the wide variety of cells that grow in the body. Rather than focus on instrumenting biological cells using electrical signals, this work makes electronic devices work more like the cells themselves. It is an approach that recognises the way in which most pharmaceutical drugs work. They interact with the proteins that are embedded in the soapy "lipid" membrane that coats each cell.
Lipid membranes are incredibly flexible although if stretched too far they will burst. But that flexibility provides the ability to have a wide variety of proteins form pores over the surface of the cell. Many of them are ion channels: specialised pincer-like proteins that open and close to drag in or expel specific ions, such as sodium or potassium. It is these ion channels that are mechanically controlled in recent experiments with ultrasound. As the particles that pass through the channels are charged, their flux can be converted to electrical signals that solid-state sensors are able to use.
From a modelling standpoint, the ion channels behave in a similar way to transistors. Some biological systems have molecular amplifiers that can apply gain: one protein signals to others in the membrane to convey much larger quantities of ions than the first protein would be able to do on its own. Artificial membranes with embedded ion channels can deal with biological signals directly rather than trying to sense electrical currents crossing biofilms.
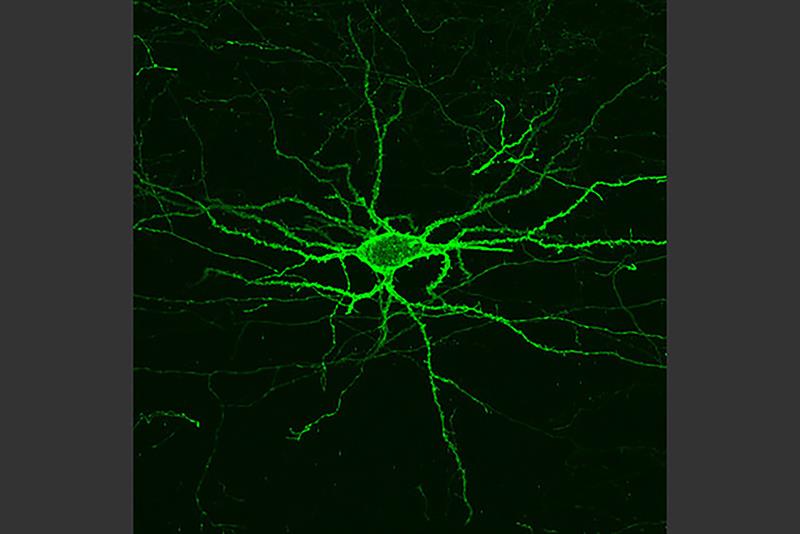 Genetic engineering carried out at MIT causes neurons to grow under fluorescent light when active
Genetic engineering carried out at MIT causes neurons to grow under fluorescent light when active
Construction is difficult as the interface between the biological membrane and the electronic systems needs to offer extremely high impedance to limit leakage as well as very low capacitance to support realistic signal bandwidths. Currents are so low that leakage is a major obstacle. However, such an interface can even be used to supply energy to an electronic system.
In a 2015 experiment, Shepard’s group used the technique to harness the cell’s own energy source, the molecule adenosine triphosphate (ATP). Proteins liberate energy stored in the bonds of this molecule by lopping off phosphate groups. Ion channels in the experimental membrane provided low levels of current to a charge pump and a group of ring oscillators.
Although biopowered sensors are attractive for smart drugs and sensor implants, applications for the membranes seem more likely to turn up in electronic systems that exploit biological proteins to increase the accuracy of sensors. For example, by embedding a variety of different ion-channel proteins in the surface, it should be possible to build taste and smell sensors that react in a similar way to those found in mammals. In principle, it would be possible to construct electronic noses that have the sensitivity of those found in dogs. However, such a device would involve finding a way to recruit the much more complex groups of ion channels found in mammals. In the short term it will be easier to target channels that identify specific molecules.
Although it has taken decades to reach this point, the recent discovery of multiple techniques for accessing the information processed by biological cells promises to speed up the process.